Data Studio Interactive Analyses
Overview
Data Studio Interactive Analyses is a public project created with the goal of enhancing end-to-end bioinformatics analysis on the Platform. The suite is currently comprised of the following interactive analyses:
- Single Cell RNA-Seq Interactive Analysis. This is an interactive analysis for performing Clustering and Cluster Marker Identification analysis on scRNA-Seq data.
- methylKit Differential Methylation Analysis. This interactive analysis is based on methylKit, an R package used for analysis and annotation of DNA methylation information obtained through high-throughput bisulfite sequencing.
- NanoStringNorm Gene Expression Analysis. This analysis is written as an Rmarkdown document and can be used for the normalization, visualization and differential expression of NanoString miRNA and RNA expression level.
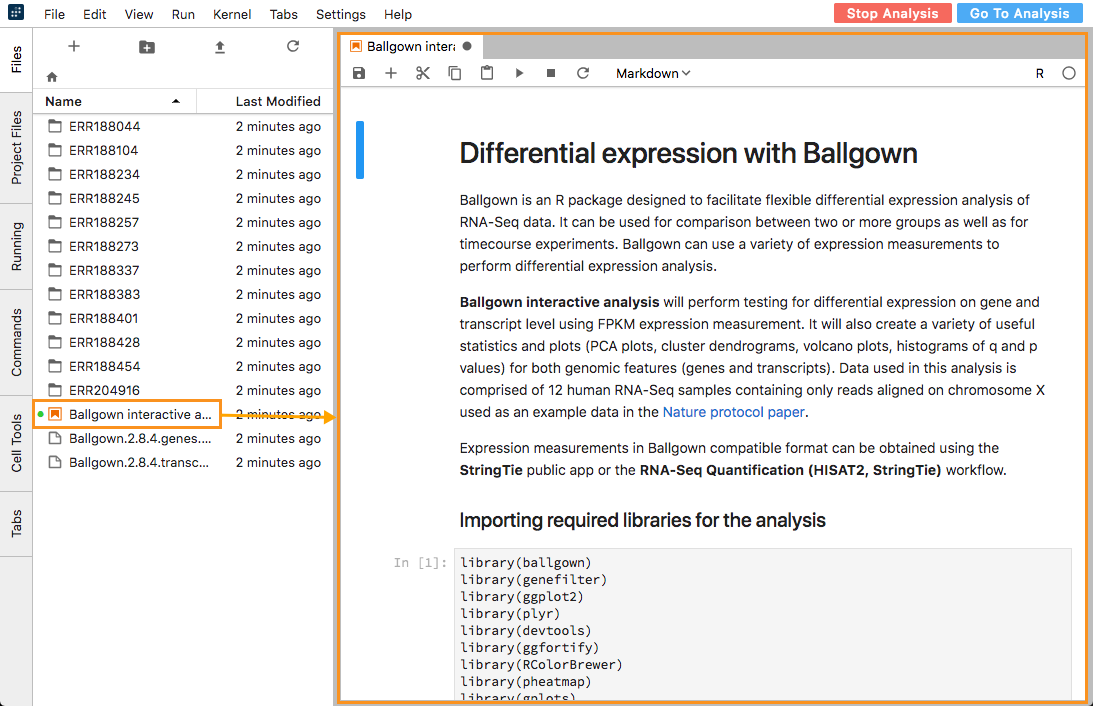
- Ballgown Interactive Analysis. This analysis tests for differential expression at the gene and transcript level using FPKM expression measurement data.
- VCF Visualization Interactive Analysis. The analysis offers several quality control analyses of variant call format (VCF) data. It accepts both raw and annotated VCF files without the need for compression or indexing.
- Structural Variation (SV) Interactive Analysis. This is a Jupyter Notebook designed for a quick overview of a VCF file containing structural variant calls. The analysis parses the SV caller’s VCF output for simple data filtering and visualization.
- ChIP-seq Interactive Analysis. The Chromatin Immunoprecipitation Sequencing (ChIP-Seq) Interactive Analysis provides a visualization of the likely locations of transcription factor binding sites or histone modifications.
- Microbiome Differential Abundance Analysis. The goal of the Microbiome Differential Abundance Interactive Analysis is to detect differential abundance of microbes between two predetermined classes of samples. This experimental design is applicable to case-control clinical studies and other settings for which there is prior assumption about the existing microbiological conditions within the different groups of samples.
For more details about each of the analyses please see our Data Studio Public Interactive Analyses blog post.
Access and run an analysis
- On the main menu bar click Public projects.
- Select Data Studio Interactive Analyses.
- Once in the public project, click the info icon next to the project title. Project information box is displayed.
- In the bottom-right part of the project information box, click Copy project.
- If needed, change the project name, url, billing group or execution settings.
- Click Copy. You are now taken to your own copy of the Data Studio Interactive Analyses public project.
- Click the Data Studio tab. You see a list of all available interactive analyses.
- Click the name of the desired analysis.
- Click Start in the top-right corner. Your analysis will start initializing.
- Once it is ready, click Open in editor in the top-right corner. This opens the JupyterLab editor.
- In the files list on the left, double click the .ipynb file containing the analysis. Your analysis is now loaded in the editor and ready to be executed.
Updated 7 months ago
